Open-ST: High-resolution spatial transcriptomics in 3D
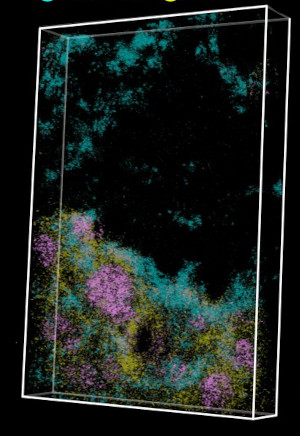
We developed a spatial transcriptomics method which is easy to use, has subcellular resolution, is cost-efficient, and scales to 3D [27], [29]. Open-ST further offers integrated H&E staining from the same tissue section and provides open-source software to process raw sequencing data, automatically align data modalities and segment cells, and align 2D sections to assemble and interactively explore 3D virtual tissue blocks.
Processing, analysis & visualization of spatial transcriptomics data
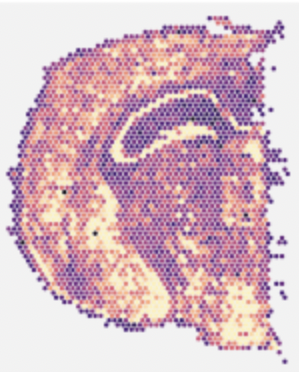
We developed a stack of tools for spatial transcriptomics datasets. Optocoder [26] uses machine learning to decode bead locations and their barcodes in array-based methods. Spacemake [24] handles any sequencing-based dataset. It offers parallel processing and additional modules: sample merging, saturation analysis, integration of spatial and scRNA-seq data. STIM [32] borrows imaging techniques to analyze and visualize large-scale spatial datasets.
Flexible spatial reconstruction of single-cell gene expression

We published an extended protocol of novoSpaRc, expanding its compatibility and showcasing its application on two new tissues: the organ of Corti and human osteosarcoma cultured cells [23]. We also made it to the front page of the current issue. (Illustration by Tulsi Voralia.)
Gene expression cartography

How to computationally reconstruct spatial positions with little or no prior knowledge from scRNA-seq data? We developed a new method (novoSpaRc) that operates under the hypothesis that nearby cells have transcriptional profiles that are often more similar than cells that are further apart [18]. We used novoSpaRc to reconstruct liver lovules, intestinal epithelium, fly and zebrafish embryos and brain cerebellum sections.
FLAM-seq: full-length mRNA sequencing
We developed a new method for high-quality sequencing of entire mRNAs, including their poly(A) tails [17]. Using human cell lines, brain organoids and C. Elegans we show that FLAM-seq delivers high-quality full-length mRNA sequences for thousands of different genes per sample.
A Single-cell transcriptome atlas of the mouse glomerulus
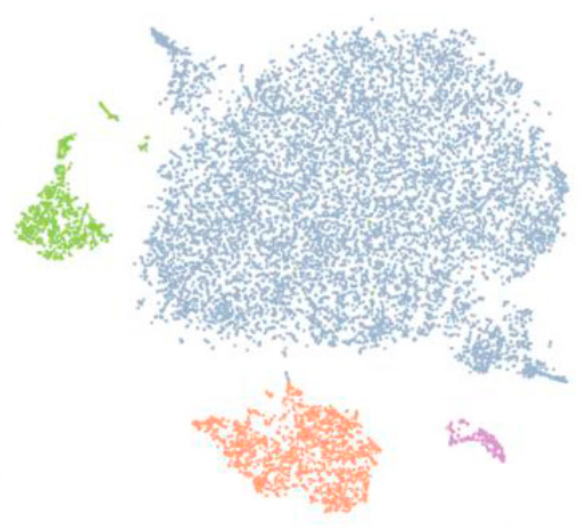
Our study comprehensively characterized gene expression in individual cells of the mouse glomerulus, setting the stage for the dissection of glomerular function at the single-cell level in health and disease [16]. We accompany our results with an online interactive atlas.
The virtual fly embryo
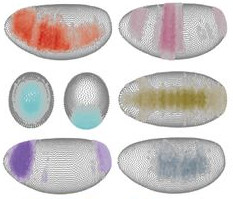
We used single-cell transcriptomics data and developed a novel computational algorithm to reconstruct a virtual Drosophila early embryo that can be queried for the expression of any given gene [15]. We also created an interactive online database that provides the transcriptional information along the virtual embryo and can be used as a resource: DVEX.
Fixing and preserving precious RNA material in single cells
It is now possible to quickly obtain the RNA of thousands of single cells at low cost. As it is critical to preserve the natural cell states, we developed a cell fixation method which stabilises the cells and preserves them over time [14]. We found our methanol-based fixation protocol to be compatible with across several different tissues and species.
Topological quanum computer elements
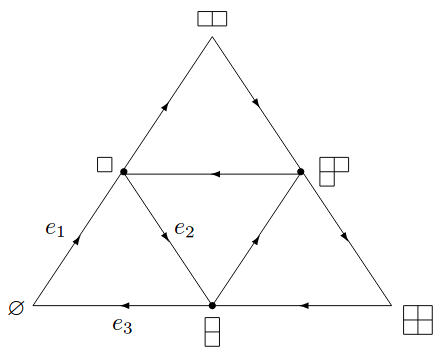
Functional quantum computers will be able to perform computations at ultra high speed and accuracy. A topological quantum computer is based on exotic quasi-particles called anyons, a family of which has been experimentally detected. We focused on extracting mathematical structures that allowed us to solve the corresponding systems exactly in [10] and [13].
Interacting fermions: a · b = - b · a
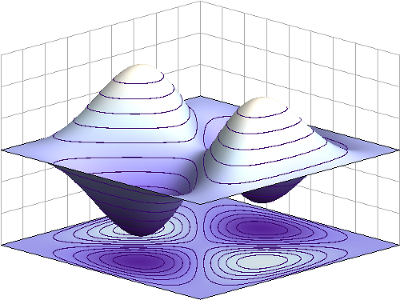
Fermions are a class of particles which obey the exclusion principle discovered by Pauli. Because of their intrinsic properties, interactions between fermions are non-trivial. We solved a 1-dimensional lattice model of interacting fermions subject to boundary conditions [8]. I then generalized the boundary conditions to a large unstudied family of supersymmetric models [9].
Impurities and defects
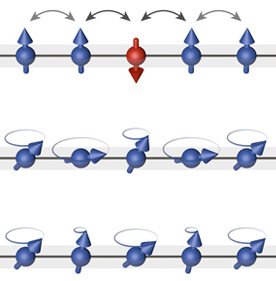
Impurities are another way to bridge the gap between theory and nature, as theoretical models are often too homogeneous to capture physical signals. We worked out the consequences of introducing impurities in different frameworks including sigma models [6], spin chains [7] and impurity boundaries [11], [12].
Life in 10 dimensions
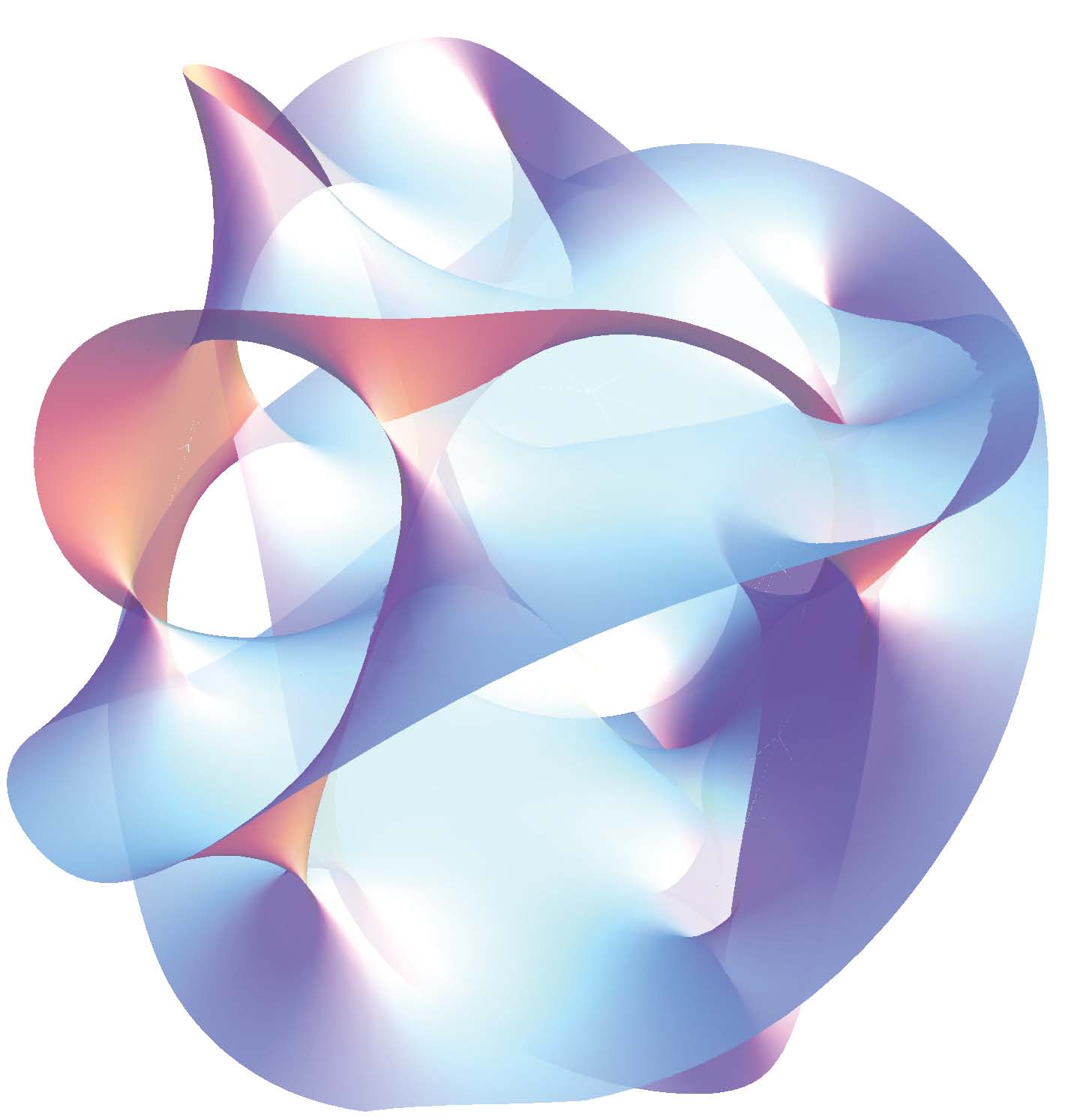

According to string theory, the world is 10-dimensional. Higher-dimensional objects exist in these spaces and interact with each other. We worked on finding new ways to embed these objects on sphere submanifolds [4].